Research
The Meyer lab applies experimental and computational strategies to measure, model, and then therapeutically manipulate cell-to-cell communication, with applications in the development of immune and cancer therapies.
Mapping and Manipulating Immune Cell Communication
Many immune receptors operate as families with multiple ligands and receptors, expressed across diverse cell populations. The lab’s efforts operate around the central hypothesis that the multiple members of these families are present to perform computation-like regulation across cell populations. Further, we can use engineering analysis tools to measure and manipulate these systems.
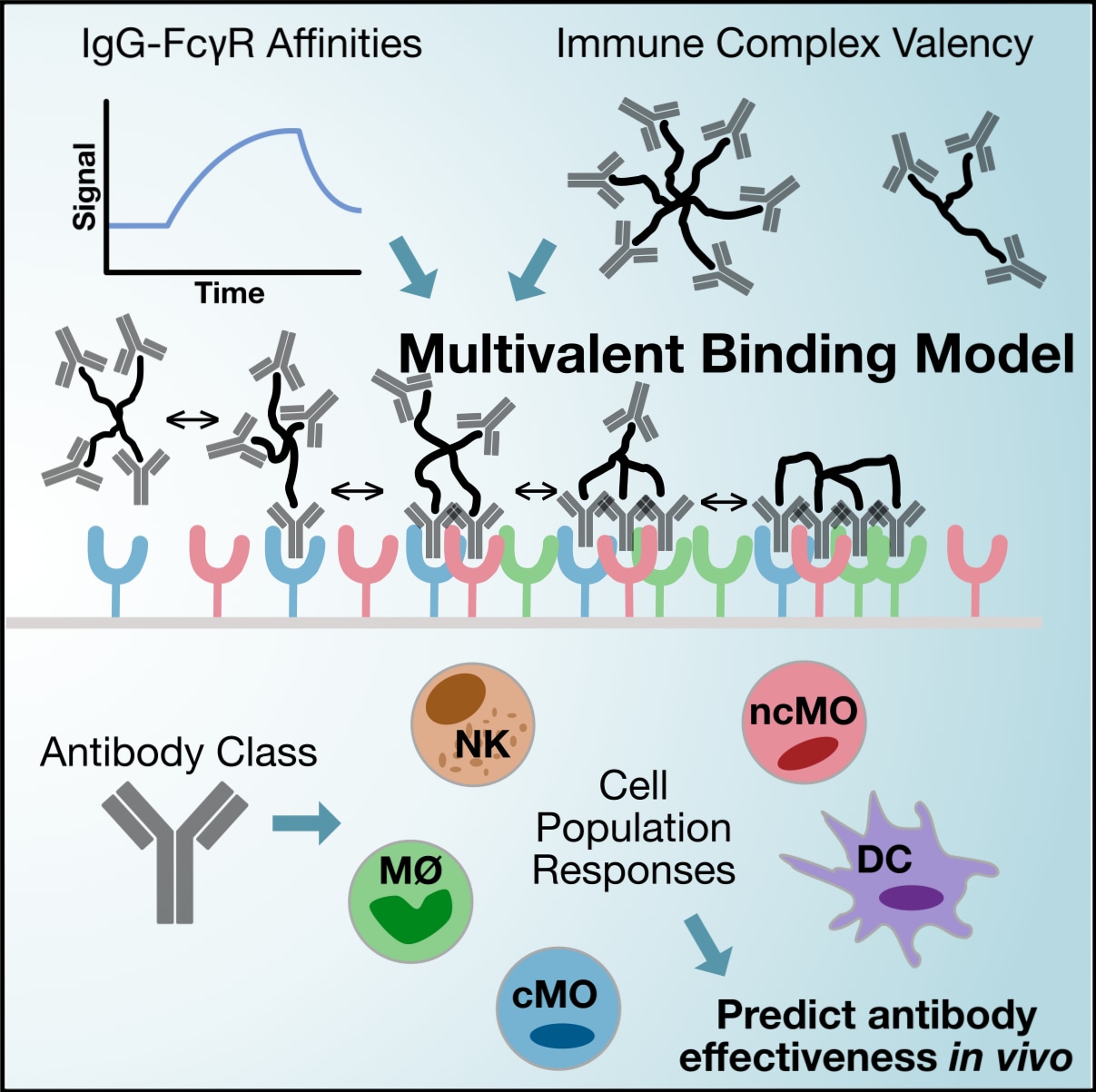
The Fc portions of IgG antibodies enable communication with many cell populations of the immune system via Fcγ receptors. The consequence of these interactions is influenced by relative affinity among the receptors, valency, and the cell populations present, creating thousands of possibilities. Using a mechanism-based binding model and data-driven analytical techniques, we aim to engineer this communication.
We are studying regulation of families like the Tyro3, AXL, MerTK (TAM) tyrosine kinases, Fcγ, and common γ-chain cytokine receptors. In addition to studying how these receptors operate as a family, we are working to develop tools that make visualizing and manipulating family-wide behavior easier.
Relevant publications: Robinett et al, Tan et al, VanDyke et al
Mapping Mechanisms of Resistance in Cancer
Targeted therapies extend many cancer patients’ lives but are limited in efficacy to a subset of patients and by the development of resistance. Efforts undertaken to identify mechanisms of resistance have uncovered numerous changes involving gene expression, post-translational regulation, and even tumor-extrinsic factors such as host-derived growth factors. Combination therapy can effectively combat resistance but requires accurate identification of the relevant resistance mechanism. Precision therapy must account for many genetic and non-genetic intrinsic and adaptive resistance mechanisms if it will accurately select these combinations.
In RTK-driven tumors, signals are transduced from the receptor to various kinases. Upon blocking the original cancer driver, resistance can be conferred by an untargeted receptor. Some receptors, however, do not provide essential resistance signals. By identifying the essential signals driving resistance from each receptor, we aim to develop measurements pinpointing the receptor causing resistance.
Projects in the lab include mapping the common essential signaling events that drive resistance, quantifying single cell heterogeneity in drug response, and exploring how the extracellular matrix environment directs resistance development.
Relevant publications: Manole et al, Claas et al, Schwartz et al, Creixell et al
Multidimensional Data Analysis
Traditional data analysis methods organize data into matrix form—a two-dimensional (2D) grid of numbers wherein each column is a measurement and each row is an observation (e.g., genes by subjects). However, this approach overlooks how measurements are often systematically collected in biology. For example, measurements to understand the molecular response of cells to therapy might be collected over concentrations of drug, time, different sources of cells, and molecular features. In these cases, the data can be organized into a multidimensional (e.g., 4D) form. Generalization of statistical tools into these multidimensional/tensor forms exist, but their use has only begun to catch on in studies of biology and medicine because there is a lack of (1) knowledge about their benefits, (2) practical and useful implementations, and (3) algorithms for specific challenges that arise with biological data. By applying these techniques, developing new algorithms, and providing accessible implementations, we are making these tools available in biomedical research.
Systems serology measurements can advance our understanding of humoral immunity. A data reduction method, “coupled matrix-tensor factorization”, effectively analyzes such data by recognizing conserved patterns and separating antigen from Fc property effects.
Relevant publications: Farhat et al, Tan et al, Orcutt-Jahns et al